Edward Tabor
Bethesda, Maryland, United States
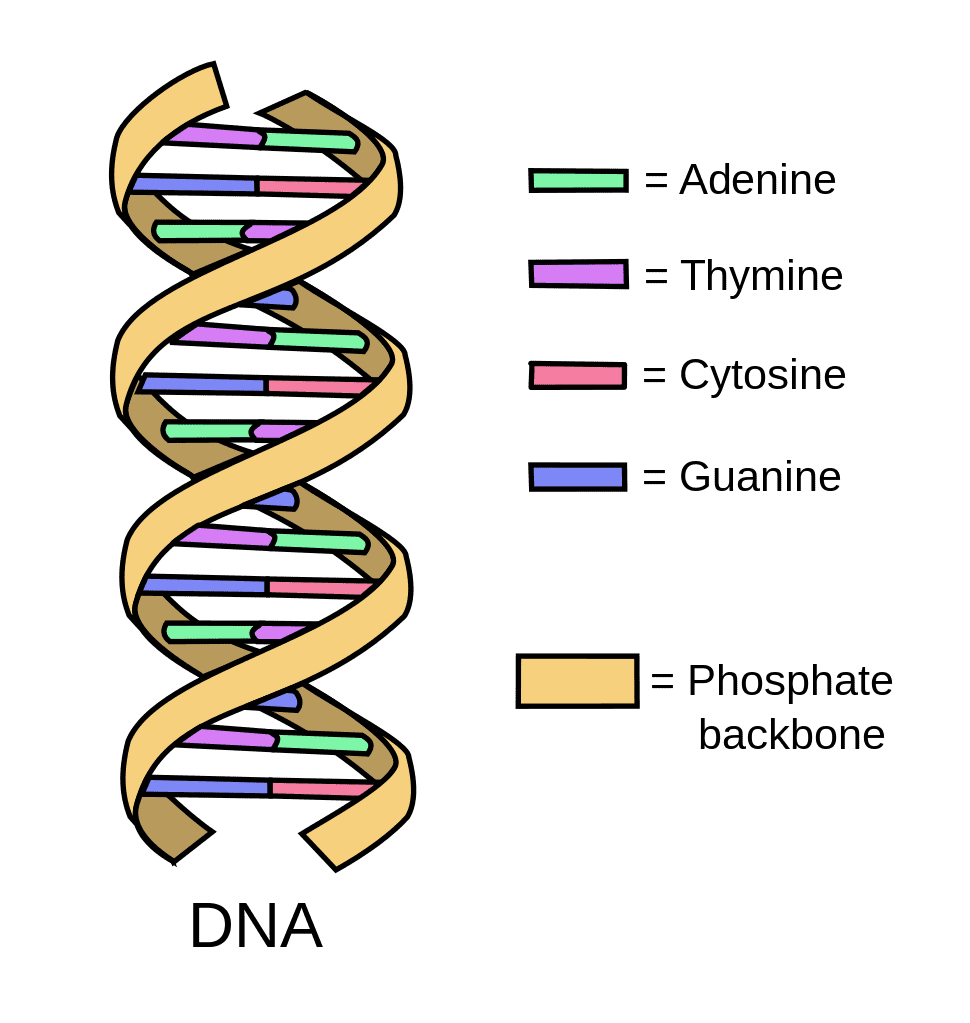
Deoxyribonucleic acid (DNA), the chemical that forms our genes, can be used to encode and transmit narrative documents and photos, as shown in several published studies. DNA might also become the next “invisible ink” because messages in DNA can be “hidden in plain sight” to reduce the chance of being detected.
Most research on DNA for storing narrative information has focused on meeting society’s future needs for archival storage. Scientists at Harvard University coded a genetics book in DNA (including 53,426 words and 11 JPG images),1 created 70 billion DNA copies of the book, and then translated it back into English. Scientists at the European Bioinformatics Institute (EBI) in the UK coded in DNA all of Shakespeare’s sonnets (in ASCII text), a color photo (in JPG format), a scientific paper (in PDF format), and a recording of a speech (in MP3 format). They created 12 million copies and sent the DNA to another laboratory where it was decoded. All of the files were back to their original condition, and the sonnets could again be read in English.2 In each of these studies, there were few coding errors,1,2 in part due to the large number of DNA copies made, which permitted checking for accuracy. DNA is ideal for this: one gram of single-stranded DNA can hold more than 215 petabytes of data,3 equal to the capacity of 300 million compact discs, but occupies a volume smaller than a poppyseed.4 A shorter message would be essentially invisible to the naked eye.
How does this work? Genetic information in living cells is coded in DNA by means of four chemicals: adenine, thiamine, guanine, and cytosine (abbreviated by the letters A, T, G, and C), arranged in groups of three. For instance, the triplet “TAC” encodes the amino acid methionine. To code a narrative message, there are 64 possible permutations of A, T, G, and C in groups of three. Twenty-six of these can be selected to code the 26 letters of the English alphabet (or any other alphabet), and the remaining unused permutations can be used to code dummy letters to make the code more difficult to break. Alternatively, the binary system of digital data storage using “0”s and “1”s can be converted to DNA coding. For instance, “A” and “C” can be used interchangeably to mean “0” and “G” and “T” interchangeably to mean “1,” as done in the Harvard study.1 Software to do this, developed by others under the aegis of a US government research program, the Molecular Information Storage program of the Intelligence Advanced Research Projects Activity (IARPA), has also been reported.4
The message is coded in the form of overlapping segments of DNA, rather than one large DNA molecule. The Harvard study used 159-nucleotide segments1 and the EBI study used 117-nucleotide segments.2 Each segment also includes coded indexing details to show where it fits into the larger message. Overlapping segments provide duplicate copies of the message and thus serve to safeguard the coding accuracy. The DNA for coding these messages can be synthesized without the use of a pre-existing strand of DNA as a template, making the process simpler than natural DNA synthesis in a cell.3
Throughout history, invisible inks, and later microdots, were used to hide coded messages. In a similar way, DNA-encoded messages “hidden in plain sight” can prevent, or at least delay, attempts by outsiders to decode them. For instance, DNA messages placed in dried saliva on the gummed seals of envelopes would be undetected unless someone knew that this contained a coded message. A smudge of DNA can be hidden on a scrap of tissue or a piece of clothing, particularly if hidden in the midst of other DNA smudges. Only the intended recipient would know where to look for it and what DNA primers to use to amplify the message.
Hiding the message requires minimal effort. DNA is hardy; it is not destroyed by long storage, cold temperatures, or mild heat, particularly if the fragments are short. Dry DNA is often readable after thousands of years in the ground.5 In addition, methods have been developed to make DNA stable even in wet conditions by creating “mirror image DNA”6 (with “L” chirality instead of the “D” chirality of natural DNA), which cannot be degraded by normal enzymes found in the environment.
Portable equipment is available to decode messages in DNA. A handheld DNA sequencer can be as small as four inches long, weighing 90 grams, and can be plugged into a laptop. Thus, DNA messages can be read almost anywhere, allowing a government agency to send messages to its agents in the field.
The use of DNA for storing non-genetic data has been reported in the scientific literature. Thus, it is likely that the methods described here have already been attempted for sending messages in secrecy. Countermeasures will no doubt be sought, such as ways to differentiate artificial DNA from natural DNA in everyday locations. This might be difficult because of the ubiquity of natural DNA in our environment, and intercepting specific messages might have to rely on information from double agents or from intercepted communications regarding the locations of DNA-encoded messages.
References
- Church GM, Gao Y, Kosuri S. “Next-generation digital information storage in DNA.” Science 337:1628, 2012.
- Goldman N, Bertone P, Chen S, et al. “Towards practical, high-capacity, low-maintenance information storage in synthesized DNA.” Nature 494:77-80, 2013.
- Hysolli E. “A DNA synthesis and decoding strategy tailored for storing and retrieving digital information.” Wyss Institute website, Harvard University. August 6, 2019. https://wyss.harvard.edu/news/save-it-in-dna/.
- Ionkov L, Settlemyer B. “DNA: The ultimate data-storage solution.” Scientific American, May 28, 2021. https://www.scientificamerican.com/article/dna-the-ultimate-data-storage-solution/.
- Reitsema LJ, Mittnik A, Kyle B, et al. “The diverse genetic origins of a Classical period Greek army.” PNAS 2022. pnas.org/doi/pdf/10.1073/pnas.2205272119.
- Fan C, Deng Q, Zhu TF. “Bioorthogonal information storage in L-DNA with a high-fidelity mirror-image Pfu DNA polymerase.” Nature Biotechnology 39:1548-1555, 2021.
EDWARD TABOR, M.D. has worked at the US Food and Drug Administration, the National Cancer Institute (NIH), and Fresenius Kabi. He has published widely on viral hepatitis, liver cancer, and pharmaceutical regulatory affairs.
Highlighted in Frontispiece Volume 15, Issue 4 – Fall 2023